Abstract
Cross-regulation of Toll-like receptor (TLR) responses by cytokines is essential for effective host defense, avoidance of toxicity and homeostasis, but the underlying mechanisms are not well understood. Our comprehensive epigenomics approach to the analysis of human macrophages showed that the proinflammatory cytokines TNF and type I interferons induced transcriptional cascades that altered chromatin states to broadly reprogram responses induced by TLR4. TNF tolerized genes encoding inflammatory molecules to prevent toxicity while preserving the induction of genes encoding antiviral and metabolic molecules. Type I interferons potentiated the inflammatory function of TNF by priming chromatin to prevent the silencing of target genes of the transcription factor NF-κB that encode inflammatory molecules. The priming of chromatin enabled robust transcriptional responses to weak upstream signals. Similar chromatin regulation occurred in human diseases. Our findings reveal that signaling crosstalk between interferons and TNF is integrated at the level of chromatin to reprogram inflammatory responses, and identify previously unknown functions and mechanisms of action of these cytokines.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Brenner, D., Blaser, H. & Mak, T.W. Regulation of tumour necrosis factor signalling: live or let die. Nat. Rev. Immunol. 15, 362–374 (2015).
Kalliolias, G.D. & Ivashkiv, L.B. TNF biology, pathogenic mechanisms and emerging therapeutic strategies. Nat. Rev. Rheumatol. 12, 49–62 (2016).
Park, S.H., Park-Min, K.H., Chen, J., Hu, X. & Ivashkiv, L.B. Tumor necrosis factor induces GSK3 kinase-mediated cross-tolerance to endotoxin in macrophages. Nat. Immunol. 12, 607–615 (2011).
Hu, X. & Ivashkiv, L.B. Cross-regulation of signaling pathways by interferon-gamma: implications for immune responses and autoimmune diseases. Immunity 31, 539–550 (2009).
Biswas, S.K. & Lopez-Collazo, E. Endotoxin tolerance: new mechanisms, molecules and clinical significance. Trends Immunol. 30, 475–487 (2009).
Ivashkiv, L.B. & Donlin, L.T. Regulation of type I interferon responses. Nat. Rev. Immunol. 14, 36–49 (2014).
Ganal, S.C. et al. Priming of natural killer cells by nonmucosal mononuclear phagocytes requires instructive signals from commensal microbiota. Immunity 37, 171–186 (2012).
Ivashkiv, L.B. Type I interferon modulation of cellular responses to cytokines and infectious pathogens: potential role in SLE pathogenesis. Autoimmunity 36, 473–479 (2003).
Prinz, M. et al. Distinct and nonredundant in vivo functions of IFNAR on myeloid cells limit autoimmunity in the central nervous system. Immunity 28, 675–686 (2008).
Stark, G.R. & Darnell, J.E. Jr. The JAK-STAT pathway at twenty. Immunity 36, 503–514 (2012).
Ramirez-Carrozzi, V.R. et al. A unifying model for the selective regulation of inducible transcription by CpG islands and nucleosome remodeling. Cell 138, 114–128 (2009).
Smale, S.T. Selective transcription in response to an inflammatory stimulus. Cell 140, 833–844 (2010).
Smale, S.T., Tarakhovsky, A. & Natoli, G. Chromatin contributions to the regulation of innate immunity. Annu. Rev. Immunol. 32, 489–511 (2014).
Smale, S.T. & Natoli, G. Transcriptional control of inflammatory responses. Cold Spring Harb. Perspect. Biol. 6, a016261 (2014).
Ivashkiv, L.B. Epigenetic regulation of macrophage polarization and function. Trends Immunol. 34, 216–223 (2013).
Heinz, S., Romanoski, C.E., Benner, C. & Glass, C.K. The selection and function of cell type-specific enhancers. Nat. Rev. Mol. Cell Biol. 16, 144–154 (2015).
Ivashkiv, L.B. & Park, S.H. Epigenetic regulation of myeloid cells. Microbiol. Spectr. 4: MCHD-0010-2015. doi:10.1128/microbiolspec.MCHD-0010-2015 (2016).
Gosselin, D. et al. Environment drives selection and function of enhancers controlling tissue-specific macrophage identities. Cell 159, 1327–1340 (2014).
Lavin, Y. et al. Tissue-resident macrophage enhancer landscapes are shaped by the local microenvironment. Cell 159, 1312–1326 (2014).
Ostuni, R. et al. Latent enhancers activated by stimulation in differentiated cells. Cell 152, 157–171 (2013).
Kaikkonen, M.U. et al. Remodeling of the enhancer landscape during macrophage activation is coupled to enhancer transcription. Mol. Cell 51, 310–325 (2013).
Saeed, S. et al. Epigenetic programming of monocyte-to-macrophage differentiation and trained innate immunity. Science 345, 1251086 (2014).
Okabe, Y. & Medzhitov, R. Tissue-specific signals control reversible program of localization and functional polarization of macrophages. Cell 157, 832–844 (2014).
Buenrostro, J.D., Giresi, P.G., Zaba, L.C., Chang, H.Y. & Greenleaf, W.J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Piper, J. et al. Wellington: a novel method for the accurate identification of digital genomic footprints from DNase-seq data. Nucleic Acids Res. 41, e201 (2013).
Foster, S.L., Hargreaves, D.C. & Medzhitov, R. Gene-specific control of inflammation by TLR-induced chromatin modifications. Nature 447, 972–978 (2007).
Yan, Q. et al. Nuclear factor-κB binding motifs specify Toll-like receptor-induced gene repression through an inducible repressosome. Proc. Natl. Acad. Sci. USA 109, 14140–14145 (2012).
Shalova, I.N. et al. Human monocytes undergo functional re-programming during sepsis mediated by hypoxia-inducible factor-1α. Immunity 42, 484–498 (2015).
Chen, J. & Ivashkiv, L.B. IFN-γ abrogates endotoxin tolerance by facilitating Toll-like receptor-induced chromatin remodeling. Proc. Natl. Acad. Sci. USA 107, 19438–19443 (2010).
Ramirez-Carrozzi, V.R. et al. Selective and antagonistic functions of SWI/SNF and Mi-2beta nucleosome remodeling complexes during an inflammatory response. Genes Dev. 20, 282–296 (2006).
Whyte, W.A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013).
Thurman, R.E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012).
Wang, H. et al. IFN-beta production by TLR4-stimulated innate immune cells is negatively regulated by GSK3-β. J. Immunol. 181, 6797–6802 (2008).
Chen, X., El Gazzar, M., Yoza, B.K. & McCall, C.E. The NF-kappaB factor RelB and histone H3 lysine methyltransferase G9a directly interact to generate epigenetic silencing in endotoxin tolerance. J. Biol. Chem. 284, 27857–27865 (2009).
Benayoun, B.A. et al. H3K4me3 breadth is linked to cell identity and transcriptional consistency. Cell 158, 673–688 (2014).
Shi, L. et al. The SLE transcriptome exhibits evidence of chronic endotoxin exposure and has widespread dysregulation of non-coding and coding RNAs. PLoS One 9, e93846 (2014).
Zhang, Z. et al. H3K4 tri-methylation breadth at transcription start sites impacts the transcriptome of systemic lupus erythematosus. Clin. Epigenetics 8, 14 (2016).
Ghisletti, S. et al. Identification and characterization of enhancers controlling the inflammatory gene expression program in macrophages. Immunity 32, 317–328 (2010).
Qiao, Y. et al. Synergistic activation of inflammatory cytokine genes by interferon-γ-induced chromatin remodeling and toll-like receptor signaling. Immunity 39, 454–469 (2013).
Glass, C.K. & Natoli, G. Molecular control of activation and priming in macrophages. Nat. Immunol. 17, 26–33 (2016).
Amit, I., Winter, D.R. & Jung, S. The role of the local environment and epigenetics in shaping macrophage identity and their effect on tissue homeostasis. Nat. Immunol. 17, 18–25 (2016).
Netea, M.G. et al. Trained immunity: A program of innate immune memory in health and disease. Science 352, aaf1098 (2016).
Lauberth, S.M. et al. H3K4me3 interactions with TAF3 regulate preinitiation complex assembly and selective gene activation. Cell 152, 1021–1036 (2013).
Sharif, M.N. et al. Twist mediates suppression of inflammation by type I interferons and Axl. J. Exp. Med. 203, 1891–1901 (2006).
Abt, M.C. et al. Commensal bacteria calibrate the activation threshold of innate antiviral immunity. Immunity 37, 158–170 (2012).
Schmidl, C., Rendeiro, A.F., Sheffield, N.C. & Bock, C. ChIPmentation: fast, robust, low-input ChIP-seq for histones and transcription factors. Nat. Methods 12, 963–965 (2015).
Donlin, L.T., Jayatilleke, A., Giannopoulou, E.G., Kalliolias, G.D. & Ivashkiv, L.B. Modulation of TNF-induced macrophage polarization by synovial fibroblasts. J. Immunol. 193, 2372–2383 (2014).
Acknowledgements
We thank A. Pernis and I. Rogatsky for discussions and review of the manuscript, and F. Barrat (Hospital for Special Surgery) for monocytes from donors with SLE. Supported by the US National Institutes of Health and The Tow Foundation (for the David Z. Rosensweig Genomics Center).
Author information
Authors and Affiliations
Contributions
S.H.P. conceived of, designed and performed most of the experiments, performed bioinformatic analysis and wrote the manuscript; Kyuho K., Y.Q., G.K., and K.-H.P.-M. contributed experiments and expertise; E.G. and Keunsoo K. performed bioinformatic analysis; L.B.I. conceived of and oversaw the study and edited the manuscript; and all authors reviewed and provided input on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Transcriptional profiles of TNF-treated macrophages, stratified by K-means clustering.
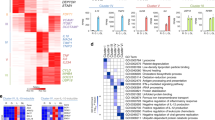
(a) This figure corresponds to Fig. 1b and shows more detail of gene expression in the different clusters within each class (right bar graphs). The left panel depicts K-means clustering (K = 12) of 1,574 LPS-induced genes (>3-fold) in the indicated conditions, showing relative gene expression for each gene. The adjacent bar graphs represent FPKM values for a given cluster in RNA-seq analysis. Error bars indicate SEM. The right two panels are bar graphs and box plots depicting FPKM values for the gene classes shown in Fig. 1b. (b) RT-qPCR analysis of representative genes in human primary macrophages stimulated with TNF (10 ng/ml) for 24 h and challenged with LPS (10 ng/ml) for 3 h; results are presented relative to baseline expression in naive macrophages (N), set as 1. Data are from one representative out of four-seven independent experiments with different blood donors with similar results. Error bars indicate SEM. (c) Induction of Class 3 genes by LPS under tolerized conditions is dependent on IFNAR signaling. RT-qPCR analysis of representative Class 3 and 4 genes in human primary macrophages stimulated with TNF (10 ng/ml) for 24 h in the presence of anti-IFNAR (5 μg/ml) or isotype CTL, and challenged with LPS (10 ng/ml) for 3 h; results are presented relative to baseline expression in unstimulated (N) cells, set as 1. Data are from one representative of three independent experiments with different blood donors with similar results. Error bars indicate SEM.
Supplementary Figure 2 Kinetic analysis of TNF-induced mRNA transcripts and TNF-induced memory.
(a) Kinetic analysis of TNF-induced mRNA transcripts. RNA-seq analysis of a time course of TNF stimulation (1, 3, 6, 24 h) was performed and the genes in the 12 clusters and 6 gene classes from Fig. 1b were visualized on a heat map (left). The gene order in each column is the same as the heat maps in Fig. 1b. Bar graphs (right) represent FPKM values for each cluster in the various gene classes. Error bars indicate SEM. (b) Venn diagram showing the overlap between LPS (10 ng/ml for 3 h) or TNF (10 ng/ml for 1 or 3 h) inducible genes, and how the genes in each region segregate into the above-defined gene classes. Genes that are not LPS-inducible are not included in the analysis in this study. (c) TNF-induced tolerization of TNF and IL6 genes persists after TNF washout. RT-qPCR analysis of TNF or IL6 genes in macrophages cultured with TNF (20 ng/ml) for 24 h, followed by washout of TNF and incubation in fresh medium for 24 or 48 h. Then, mRNA levels were measured after LPS (10 ng/ml for 3 h) stimulation. Data are from one representative of three independent experiments with different blood donors with similar results. Error bars indicate SEM.
Supplementary Figure 3 Class-specific chromatin regulation in TNF-treated macrophages.
(a) Quantitation of the data shown as a heat map in Fig. 2a. Box graphs represent H4ac, H3K4me3, H2Bub ChIP-seq and ATAC-seq normalized tag densities (log2) at the promoters (-2kb < TSS < +2kb) of a given class under the indicated conditions. **** p < 0.0001, ** p < 0.01, * p < 0.05, Kolmogorov-Smirnov test. (b) Representative UCSC Genome Browser tracks displaying normalized profiles for H4ac, H3K4me3, H2Bub ChIP-seq and ATAC-seq signals at Class 1(TNF, IL27, TNFSF9 and TNFAIP2), Class 3 (OASL, IFIT1, IRF7 and MX1) and Class 4 (FPR1/2, PTGES, CH25H, CLU) genes under indicated conditions (N, L, T, T-L). Boxes enclose genomic regions regulated across conditions. (c) Box graphs represent quantitation of the normalized tag densities (log2) for 6 histone modifications and chromatin accessibility at the promoters (-2kb < TSS < +2kb) in naïve macrophages. Each class we tested is indicated in x-axis. **** p < 0.0001, *** p < 0.001, ** p < 0.01, * p < 0.05, Kolmogorov-Smirnov test. (d) Heatmaps of H3K36me3, H3K56ac, and H3K79me2 normalized tag densities at gene bodies (TSS to TES) of a given class under indicated conditions. (e) Tolerization with TNF does not increase negative H3K27me3 or H3K9me3 histone marks at Class 1 and 2 tolerized gene promoters (-2kb < TSS < +2kb). Relative ChIp-seq normalized tag counts (upper) and ChIP-qPCR confirmation at IL6 promoter (lower).
Supplementary Figure 4 Class-specific chromatin regulation.
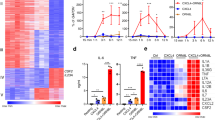
(a) Box graphs represent quantification of the normalized tag densities for H2Bub, H3K4me3 ChIP-seq and ATAC-seq at the promoters (-2kb < TSS < +2kb) in L and T-L conditions. ATAC-seq data represent pooled samples from three (T-L) or five (L) replicates from independent donors, and H2Bub, H3K4me3 ChIP-seq data are from an additional independent donor. p-value, Kolmogorov-Smirnov test. (b) Formaldehyde-assisted isolation of regulatory elements (FAIRE) assay at representative TNF, IL6 and CCL5 genes under indicated conditions. Data from two experiments out of four to six independent experiments with different donors are shown. The decrease in chromatin accessibility in T-L versus L conditions (condition 4 versus 2) at TNF and IL6 promoters was consistently observed. A decrease in basal chromatin accessibility at TNF and IL6 promoters after TNF treatment (condition 3 versus 1) was not observed in all donors, and data representative of the two patterns observed are shown separately as donor 1 (left) and donor 2 (right). (c) Distribution of ATAC-seq footprints and occupancy of transcription factor binding sites. Heatmaps showing per nucleotide ATAC-seq cleavage sites for NRF, NFY and SP1 motifs in LPS-stimulated human primary macrophages ranked by tag density. The number of ATAC-seq footprints for each TF is shown in y-axis. 200 bp windows are shown centered at the midpoints of the ATAC-seq footprint. Footprinting was performed with two independent ATAC-seq replicates. (d-f) Reversal of tolerization of IL6 by GSK3 inhibition is mediated by increased type I IFN. (d) Inhibition of GSK3 results in increased IFNB expression after TNF stimulation. RT-qPCR analysis of IFNB mRNA in human primary macrophages stimulated with TNF (10 ng/ml) for indicated times in the presence of SB216763 (10 μM) or control DMSO. (e) Increased Ifnb expression in GSK3-deficient macrophages. RT-qPCR analysis of Ifnb mRNA in BMDMs obtained from mice lacking myeloid GSK3 (Gsk3bfl/fl Cre) or genetically matched control mice (Gsk3b+/+ Cre), then stimulated with TNF (40 ng/ml) for indicated times (left) or for 24 h and then challenged for 3 h with LPS (10 ng/ml, right). (f) IFNAR blockade prevents reversal of tolerance by GSK3 inhibitor SB216763. RT-qPCR analysis of IL6 mRNA in human primary macrophages stimulated with TNF (10 ng/ml) for 24 h and then challenged with LPS (10 ng/ml) for 3 h in the presence of SB216763 (10 μM), anti-IFNAR (5 μg/ml) and isotype CTL as indicated. (g) RT-qPCR analysis of TNF and IL-6 mRNA in human primary macrophages under indicated conditions. IFN-α reversed TNF-induced tolerance (bar 12) but did not reverse LPS-induced endotoxin tolerance (bar 10). mRNA results are presented relative to baseline expression in unstimulated cells, set as 1 (d-g). Data are representative of three independent experiments (d, f, g) or two independent experiments (e).
Supplementary Figure 5 Enhancer profiles of TNF-treated macrophages.
(a) Heatmaps of H3K27ac and ATAC-seq normalized tag densities at the enhancers (see Methods) of genes in different classes based on Fig. 1b (upper). Box graphs represent quantitation of the normalized tag densities (log2) for a given class (lower). **** p < 0.0001, ** p < 0.01, * p < 0.05, Kolmogorov-Smirnov test. (b) The number of genes (%) that are associated with super enhancers and typical enhancers in a given gene class in the indicated conditions. (c-d) Bar graphs represent the number of genes and % of genes in each class that are associated with latent enhancers (c) or CpG islands (d). (e) Significantly enriched motifs (p < 10-5) in ATAC-seq footprints at enhancers in each gene class (see Methods) in indicated conditions. Motifs are grouped according to transcription factor families. (f) De novo motif analysis of enhancers (entire ATAC-seq peak in all experimental conditions) associated with the 6 gene classes using HOMER.
Supplementary Figure 6 Regulation of chromatin accessibility by TNF and IFN-α.
(a) Correlation matrix heatmap based on Pearson correlation coefficients comparing normalized RNA-seq tag counts across all indicated conditions and replicates. The concordance between replicates was very high (the number in each box indicates Pearson correlation coefficient, r). (b) Heatmap of ATAC-seq, and H2Hub and H3K4me3 ChIP-seq normalized tag densities of Class 1 genes (upper) under all eight experimental conditions performed together in one experiment. Gene order is the same as in Fig. 1b. Box graphs represent quantitation of the normalized tag densities (log2) for Class 1 gene promoters (lower). p-value, Kolmogorov-Smirnov test. (c) Heatmap of ATAC-seq normalized tag densities at enhancers (see Methods) of Class 1 genes in indicated conditions (upper). Box graphs represent quantitation of the normalized tag densities (log2) for Class 1 gene enhancers (lower). **** p < 0.0001, Kolmogorov-Smirnov test. (d) Representative UCSC Genome Browser tracks displaying normalized profiles for ATAC-seq and H2Bub, H3K4me3 ChIP-seq signals at IL27, TNFSF9, SPHK1, IL6, CCL20 and CXCL2 Class 1 genes under indicated conditions. Boxes enclose genomic regions regulated across conditions.
Supplementary Figure 7 Co-occupancy of IRF1 and p65 at genomic regions associated with class 1 genes encoding inflammatory molecules.
(a) De novo motif enrichment analysis of Class 1 gene enhancers using ATAC-seq footprints under same conditions as the promoter analysis in Fig. 6a. (b) Bar graphs represent cumulative values for IRF family members in RNA-seq analysis from three replicates. (c) Representative UCSC Genome Browser tracks displaying normalized profiles for p65 and IRF1 ChIP-seq signals at CXCL10, CXCL11, IL8, TNF, TNIP2, CCL20 and IL6 (Class 1) genes under indicated conditions. Boxes enclose co-localization of p65 and IRF1 binding peaks to the same genomic regions. (d) ChIP-qPCR analysis of recruitment of p65 and IRF1 to representative genes under indicated conditions. Data are representative of three independent experiments. (e-f) Venn diagram showing the overlap between p65 ChIP-seq peaks in human macrophages (stimulated with TNF (10 ng/ml) for 24 h) and IRF1 Chip-seq peaks (stimulated with IFN-γ (100 U/ml) for 24 h and challenged with LPS (10 ng/ml) for 3 h) within 10 kb ± TSS of Class 1 genes. p-value, Fisher’s exact test relative to a randomly selected control gene set. Representative UCSC Genome Browser tracks displaying normalized profiles for p65 and IRF1 ChIP-seq signals at IL27 and IL1B genes. Boxes enclose co-localization of p65 and IRF1 binding peaks to the same genomic regions. IRF1 ChIP-seq data are from GSE43036.
Supplementary Figure 8 ‘Sequential rheostat’ model of integrated regulation of the TLR-induced transcriptional response by TNF and IFN-α.
(a) Sequential rheostat model of integrated regulation of the TLR-induced transcriptional response by TNF and IFN-α via upstream signaling and downstream chromatin-based epigenetic mechanisms. Regulation of TLR-induced signaling pathways and chromatin states of TLR-inducible genes by TNF and IFN-α determines transcriptional output. TNF treatment ‘tolerizes’ macrophages to diminish signaling responses to TLR4 agonist LPS. Left panel. In TNF-stimulated cells, IFN-α promotes opening of chromatin at tolerized (T) genes, thereby allowing gene induction in response to weak upstream signals. Right panel. At nontolerized (NT) genes, TNF increases chromatin accessibility concomitantly with suppression of upstream signaling. This enables NT gene induction in response to weak TLR4 signaling upon secondary LPS challenge. (b) Left panel. In naïve macrophages LPS activates a strong signal that is sufficient to broadly remodel chromatin at target genes promoters, resulting in expression of the full set of TLR-inducible genes. Right panel. TNF induces chromatin remodeling at subsets of TLR-inducible genes in a class-specific manner. In parallel, TNF induces expression of multiple signaling inhibitors such that LPS activates only a weak signal. This weak signal is insufficient to induce chromatin remodeling required for transcriptional activation of Class 1 and 2 T genes, but is sufficient to activate transcription at Class 3 and 4 NT genes where chromatin has been opened and primed by TNF. Exogenous type I IFN cooperates with TNF to open and prime chromatin at Class 1 and 2 genes, enabling transcriptional activation by weak LPS-induced signals. Thus, environmental signals are integrated at the epigenomic level to regulate the TLR response in a gene class-specific manner.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8, Supplementary Table 1, and Supplementary Notes 1 and 2.
Supplementary Table 1
NI-A23840B-Copy of 29640_1_supp_348174_brhb4p.
Rights and permissions
About this article
Cite this article
Park, S., Kang, K., Giannopoulou, E. et al. Type I interferons and the cytokine TNF cooperatively reprogram the macrophage epigenome to promote inflammatory activation. Nat Immunol 18, 1104–1116 (2017). https://doi.org/10.1038/ni.3818
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3818
This article is cited by
-
BHLHE40/41 regulate microglia and peripheral macrophage responses associated with Alzheimer’s disease and other disorders of lipid-rich tissues
Nature Communications (2024)
-
Role of IFN-α in Rheumatoid Arthritis
Current Rheumatology Reports (2024)
-
The crucial regulatory role of type I interferon in inflammatory diseases
Cell & Bioscience (2023)
-
Cancer cells resistant to immune checkpoint blockade acquire interferon-associated epigenetic memory to sustain T cell dysfunction
Nature Cancer (2023)
-
Corticosteroids reduce pathologic interferon responses by downregulating STAT1 in patients with high-risk COVID-19
Experimental & Molecular Medicine (2023)